
Image: Chinnapong

Image: Chinnapong
Research project Approximately half of patients dying from prostate cancer die despite being diagnosed before the cancer had spread, thus being potentially curable. Today, treatments for this high-risk group are obviously inadequate, but by introducing more efficient therapy in an early state, metastatic spread could be prevented and lives saved. Unfortunately, efficient treatments often come with side-effects, and as not all patients benefit from the therapy, methods for accurate treatment selection are needed.
SPRINTR aims to implement optimized biomarker-based prediction models for precision medicine in high-risk prostate cancer and thereby increase survival while decreasing unnecessary side-effects and costs for society. The project will also initiate a structure for clinical studies, which by integration of already implemented systems for clinical information with national registers and database structures can create a platform for efficient and open research for cancer precision medicine.
BACKGROUND
High-risk non-metastatic prostate cancer remains a significant cause of mortality and current treatment options are inadequate. A large proportion of men will develop progression after therapy with curative intention. In this stage, before clinically detectable metastases, we believe that intensified therapy could increase survival.
In metastatic prostate cancer, androgen deprivation therapy followed by chemotherapy and newer androgen receptor pathway-targeting therapies have improved survival. However, they also come with elevated toxicity. Precision medicine is the key to identifying patients who could benefit from therapy escalation earlier in the disease course.
Various biomarker panels, including liquid biopsies and imaging, have been proposed for patient stratification. This study will establish a precision medicine platform centered on molecular profiling, imaging, and histology to enhance survival rates for high-risk prostate cancer patients. Our plan involves analyzing retrospective cohorts to identify biomarkers, conducting a multi-center prospective clinical study to validate biomarker algorithms, and further establishing a biomarker-driven adaptive multi-center, multi-arm randomized clinical trial platform. Additionally, we will initiate a drug discovery development program based on targetable pathways identified via continuous profiling of trial participants with poor response to therapy.
GOAL
To increase survival for patients with locally advanced, high-risk PC, by establishing a precision medicine platform based on molecular profiling, imaging, and histology.
Specific aims are to:
Overall, the project's novelty is defined by its focus on high-risk PC before clinically manifest metastases, where previously no biomarkers have been implemented, and the inclusion of not only genetic but phenotypic and image-based biomarkers compared to the Probio and STAMPEDE trials. In addition, inclusion of metastasis-derived profiling algorithms increases the potential to identify patients with metastasis-prone tumors.
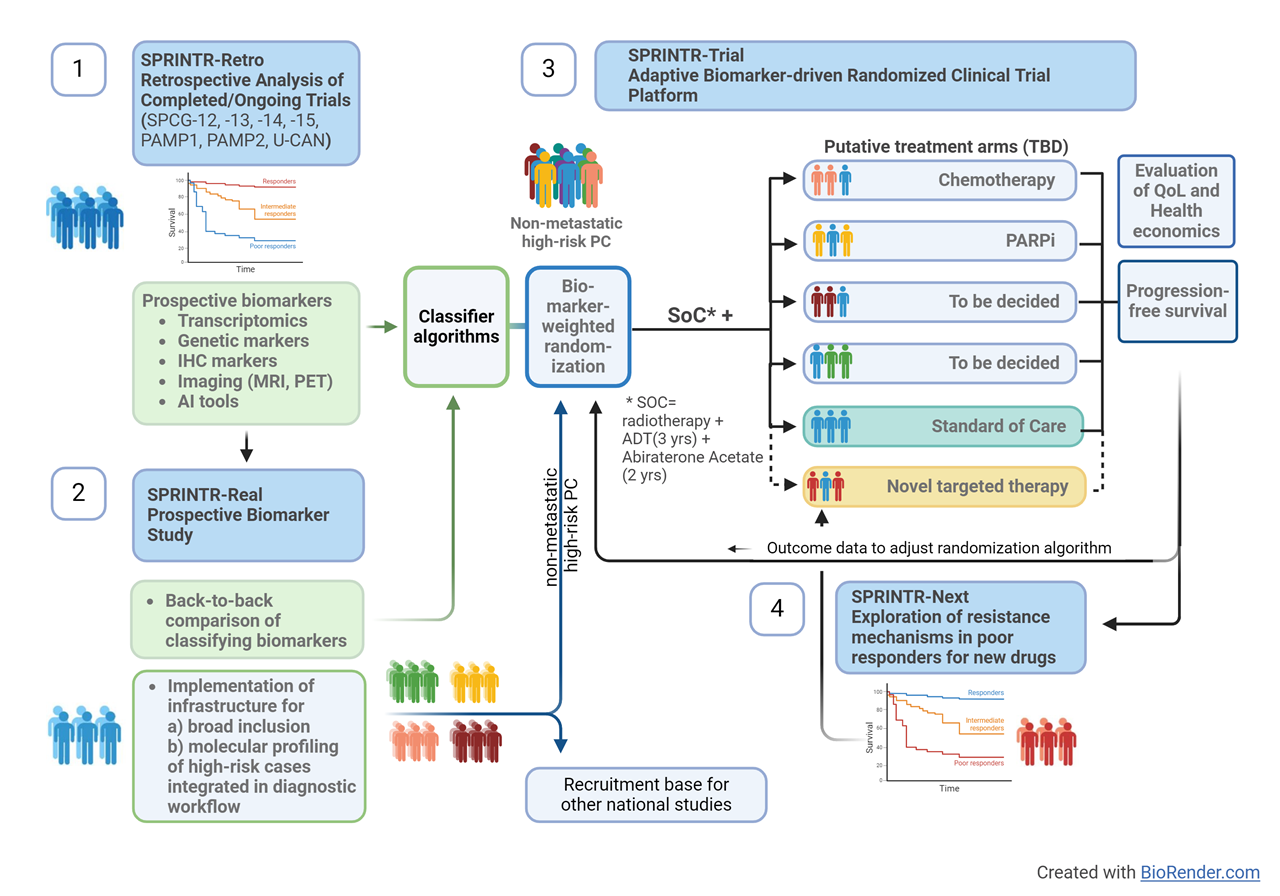
In addition, SPRINTR aims to facilitate high-quality research in the prostate cancer field in Sweden by integrating broad patient recruitment and biomarker profiling in the diagnostic workflow. Together with an automated clinical follow-up function from Swedish registers this will create a continuously growing research database as the foundation for patient selection and follow-up as a basis for other prostate cancer studies in Sweden. This will increase the possibility for high-quality, fully powered, trials with efficient recruitment, decrease the administrative burden for study personnel, and enable good and equal access to clinical trials for patients regardless of location.
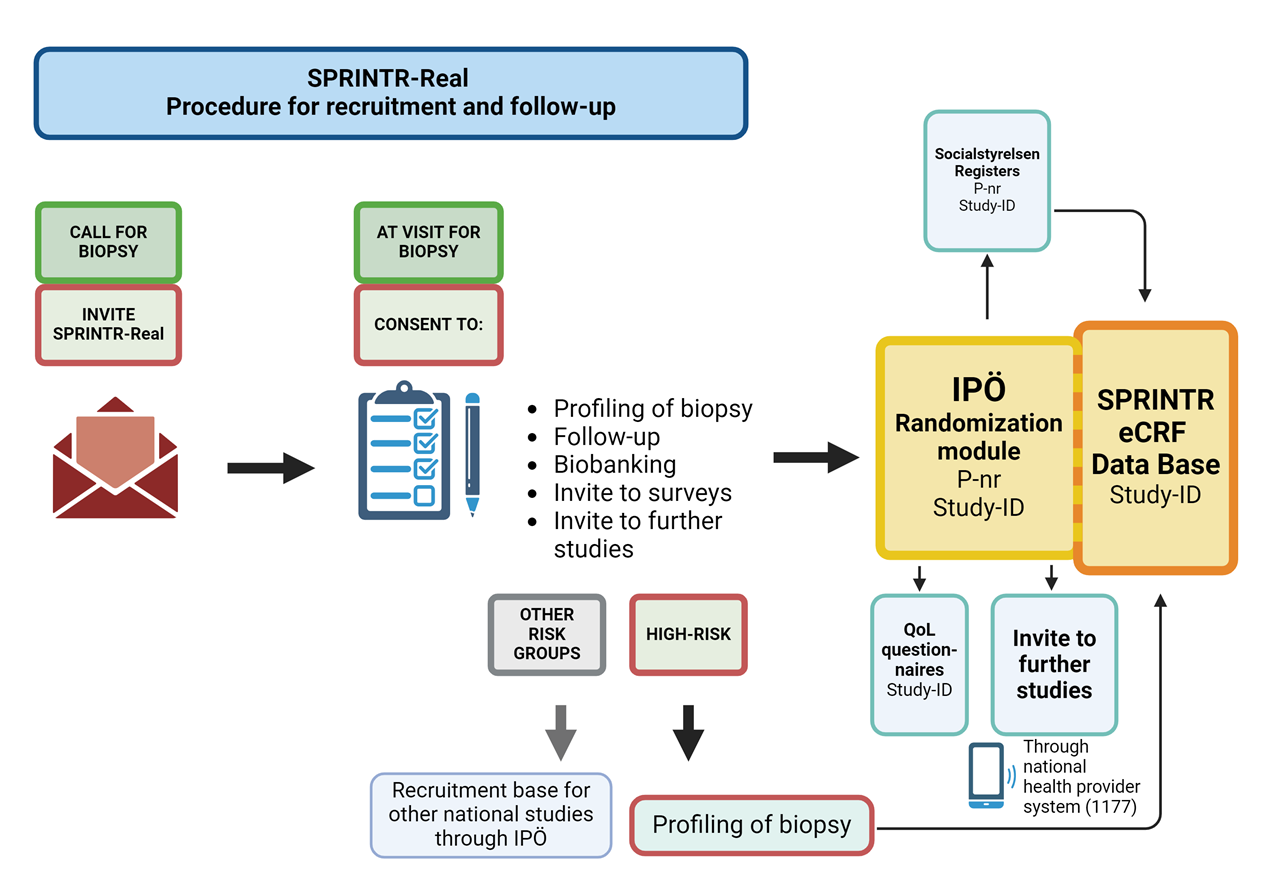
RELATED STUDIES
PARALLEL TRIALS
PUBLICATIONS
Josefsson A, Effect of docetaxel added to bicalutamide in Hormone-Naïve non-metastatic prostate cancer with rising PSA, a randomized clinical trial (SPCG-14) https://pubmed.ncbi.nlm.nih.gov/37073813/
Spyratou V, Ki67 and prostate specific antigen are prognostic in metastatic hormone naïve prostate cancer https://pubmed.ncbi.nlm.nih.gov/37713321/
Sandgren K, Histopathology-validated lesion detection rates of clinically significant prostate cancer with mpMRI, [68Ga]PSMA-11-PET and [11C]Acetate-PET https://pubmed.ncbi.nlm.nih.gov/37615497/
Wikström P, Epithelial and Stromal Characteristics of Primary Tumors Predict the Bone Metastatic Subtype of Prostate Cancer and Patient Survival after Androgen-Deprivation Therapy https://pubmed.ncbi.nlm.nih.gov/37615497/
Thysell E, Clinical and biological relevance of the transcriptomic-based prostate cancer metastasis subtypes MetA-C https://pubmed.ncbi.nlm.nih.gov/34889043/
Järemo H, Investigating microRNA Profiles in Prostate Cancer Bone Metastases and Functional Effects of microRNA-23c and microRNA-4328 https://pubmed.ncbi.nlm.nih.gov/37173903/
Josefsson A, Circulating tumor cells mirror bone metastatic phenotype in prostate cancer https://pubmed.ncbi.nlm.nih.gov/30034626/
Josefsson A, Circulating Tumor Cells as a Marker for Progression-free Survival in Metastatic Castration-naïve Prostate Cancer https://pubmed.ncbi.nlm.nih.gov/28295408/
Gonora M, Characteristics of Patients in SPCG-15-A Randomized Trial Comparing Radical Prostatectomy with Primary Radiotherapy plus Androgen Deprivation Therapy in Men with Locally Advanced Prostate Cancer https://pubmed.ncbi.nlm.nih.gov/35813256/
Björklund J, The 90-day cause-specific mortality after radical prostatectomy: a nationwide population-based study https://pubmed.ncbi.nlm.nih.gov/34191407/
Schostak M, Practical Guidance on Establishing a Molecular Testing Pathway for Alterations in Homologous Recombination Repair Genes in Clinical Practice for Patients with Metastatic Prostate Cancer https://pubmed.ncbi.nlm.nih.gov/37714762/
Merseburger AS, Apalutamide plus androgen deprivation therapy in clinical subgroups of patients with metastatic castration-sensitive prostate cancer: A subgroup analysis of the randomised clinical TITAN study https://pubmed.ncbi.nlm.nih.gov/37708629/
Thysell E, Gene expression profiles define molecular subtypes of prostate cancer bone metastases with different outcomes and morphology traceable back to the primary tumor https://pubmed.ncbi.nlm.nih.gov/31162796/
Hammarsten P, Immunoreactivity for prostate specific antigen and Ki67 differentiates subgroups of prostate cancer related to outcome https://pubmed.ncbi.nlm.nih.gov/30980038/
Lindgren Belal S, Applications of Artificial Intelligence in PSMA PET/CT for Prostate Cancer Imaging https://pubmed.ncbi.nlm.nih.gov/37357026/
Trägårdh E, Freely Available, Fully Automated AI-Based Analysis of Primary Tumour and Metastases of Prostate Cancer in Whole-Body [18F]-PSMA-1007 PET-CT https://pubmed.ncbi.nlm.nih.gov/36140502/
Björeland U, Impact of neoadjuvant androgen deprivation therapy on magnetic resonance imaging features in prostate cancer before radiotherapy https://pubmed.ncbi.nlm.nih.gov/33898790/
Sandgren K, Registration of histopathology to magnetic resonance imaging of prostate cancer https://pubmed.ncbi.nlm.nih.gov/34258403/
Chappidi, Transcriptomic Heterogeneity of Expansile Cribriform and Other Gleason Pattern 4 Prostate Cancer Subtypes https://pubmed.ncbi.nlm.nih.gov/37474400/
Das R, An integrated functional and clinical genomics approach reveals genes driving aggressive metastatic prostate cancer https://pubmed.ncbi.nlm.nih.gov/34326322/
Helzer KT Fragmentomic analysis of circulating tumor DNA-targeted cancer panels https://pubmed.ncbi.nlm.nih.gov/37330052/
de Jong AC, Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning https://pubmed.ncbi.nlm.nih.gov/37031196/
ADVISORY BOARD
Prof. N James (ICR, London, PI of STAMPEDE)
Prof. H Grönberg (KI PI ProBio)
Prof. D Larsson (GU, Head of Clinical Trials Sweden)
Prof. F Feng (UCSF, USA)
Prof T. Helleday (KI)
E Hallersjö Hult (Vision Zero Cancer)
Prof. P Cornford (UK, EAU Guidelines)
H Joelsson (Patient representative, prostatacancerförbundet)
R Rosenquist Brandell (GMS)
E Axén (NPCR registry)

A project for more efficient treatment for aggressive prostate cancer gets SEK 21 million in funding,