Opportunities
My group is currently searching for new group members, if you would like to know more please do not hesitate to contact me, silvia.remeseiro@umu.se
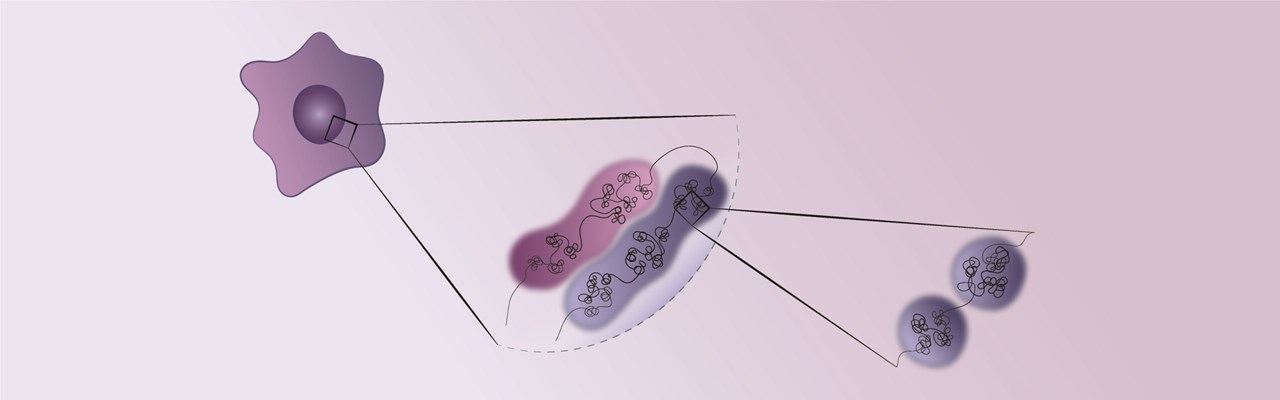
Image: Silvia Remeseiro
Research group 3D Genomics and Chromatin Biology in Brain Cancer
Remeseiro's research group focuses on 3D genomics and chromatin biology in brain cancer.
We are a multidisciplinary team combining expertise in multi-omics approaches—including 3D genomics— with advanced computational analysis, all grounded in a functional genomics framework that integrates both in vitro and in vivo validation systems. The overarching goal of our work is translational: to identify and target epigenetic regulators in order to disrupt neuron-to-glioma synaptic communication in glioblastoma.
Research interest
Glioblastoma is the most lethal and aggressive, but also the most common, of all primary brain tumors. Most genetic variants that predispose to cancer are located in non-coding regions enriched in putative enhancers, whose systematic rearrangements have a high impact on gene expression in cancer. Despite the number of susceptibility loci identified in glioblastoma, a functional understanding of how the non-coding regulatory genome contributes to the pathogenesis is missing.
Our research is focused in understanding how the reprogramming of regulatory regions and topological changes in 3D chromatin organization determine gene dysregulation in glioblastoma. We bring knowledge from different biological and technical backgrounds into the glioblastoma field, employing next generation sequencing approaches, genome editing and live-cell imaging in both glioblastoma mouse models and patient-derived glioblastoma cell lines.
The lab is currently studying the 3D chromatin organization of the glioblastoma genome and its interplay with the epigenetic and transcriptional landscapes, and how this interplay influences malignancy and integrates cues from the neurons in the tumor microenvironment to orchestrate gene expression in glioblastoma.
Strengths in the lab
In the lab we combine state-of-the-art approaches in different fields. We have a strong expertise in "omics"-based approaches (e.g HiChIP, 4C-seq, HiC, ChIP-seq, ATAC-seq, RNA-seq), bioinformatics, CRISPR-Cas9-based (epi-)genome editing, mouse genetics, mouse models of cancer and live-cell imaging. We apply these various technologies to the study of 3D chromatin organization and long-range gene regulation in glioblastoma, using both mouse models and patient-derived glioblastoma cell lines.
Publications
Pre-Prints
- Nordin A, Chakraborty C, Jonasson M, Dana O, Zambanini G, Pagella P, Silvia Remeseiro#, Cantù C#. Wnt signaling alters CTCF binding paterns and global chromatin structure. bioRxiv https://doi.org/10.1101/2024.01.22.576696
- Holm E*, Vermeulen I*, Parween S, López-Pérez A, Cillero-Pastor B, Vandenbosch M, Silvia Remeseiro+, Andreas Hörnblad#. AMPK-activator ATX-304 reduces oxidative stress and improves MASLD via metabolic switching. bioRxiv https://doi.org/10.1101/2024.02.13.578901
Published Articles
- Dennhag N, Kahsay A, Nissen I, Nord H, Chermenina M, Liu J, Arner A, Liu J, Backman L, Silvia Remeseiro+, von Hofsten J# and Pedrosa F#. fhl2b mediates extraocular muscle protection in zebrafish models of muscular dystrophies and its ectopic expression ameliorates affected body muscles. Nature Communications (2024) 15, 1950.
- Vincent C and Silvia Remeseiro#. The immune response behind peptide vaccination in diffuse midline glioma. Molecular Oncology. (2024), doi:10.1002/1878-0261.13686.
- Chakraborty C, Nissen, Vincent CA, Hägglund AC, Hörnblad A, Silvia Remeseiro#. Rewiring of the promoter-enhancer interactome and regulatory landscape in glioblastoma orchestrates gene expression underlying neurogliomal synaptic communication. Nature Communications (2023) 14, 6446.
- Vincent CA, Nissen I, Dakhel S, Hörnblad A, Silvia Remeseiro#. Epigenomic perturbation of novel EGFR enhancers reduces the proliferative and invasive capacity of glioblastoma and increases sensitivity to temozolomide. BMC Cancer (2023) 23:945.
- López-Pérez A, Silvia Remeseiro+, Hörnblad A#. Diet-induced rewiring of the Wnt gene regulatory network connects aberrant splicing to fatty liver and liver cancer in DIAMOND mice. Scientific Reports (2023) 13:18666.
- Andreas Hörnblad# and Silvia Remeseiro#. “Epigenetics, enhancer function and 3D chromatin organization during reprogramming to pluripotency.” Cells (2022) 11, 1404
- López-Pérez A, Norlin S, Steneberg P, Silvia Remeseiro+, Edlund H#, Hörnblad A#. PAN-AMPK activator O304 prevents gene expression changes and re-mobilization of histone marks in islets of diet-induced obese mice. Scientific Reports (2021) 11:24410.
- Dakhel S, Davies WIL, Joseph JV, Tomar T, Silvia Remeseiro+, Gunhaga L#. Chick fetal organ spheroids as a model to study development and disease. BMC Molecular and Cell Biology, (2021) 22, 37-17. http://doi.org/10.1186/s12860-021-00374-6
- Bahr C*, von Paleske L*, Uslu VV*, Silvia Remeseiro, Takayama N, Ng SW, Murison A, Langenfeld K, Petretich M, Scognamiglio R, Zeisberger P, Benk AS, Amit I, Zandstra PW, Lupien M, Dick JE, Trumpp A, Spitz F. A Myc enhancer cluster regulates normal and leukaemic haemotopoietic stem cell hierarchies. Nature (2018) 553:515-520.
- Symmons O, Pan L, Silvia Remeseiro, Aktas T, Klein F, Huber W, Spitz F. The Shh Topological Domain Facilitates the Action of Remote Enhancers by Reducing the Effects of Genomic Distances. Developmental Cell (2016) 39(5):529-543.
- Silvia Remeseiro*, Andreas Hörnblad*, François Spitz. Gene regulation during development in the light of topologically associating domains. Wiley Interdisciplinary Reviews Developmental Biology (2016). 5(2):169-85.
- Cuadrado A*, Silvia Remeseiro*, Graña O, Pisano DG, Losada A. The contribution of cohesin-SA1 to gene expression and chromatin architecture in two murine tissues. Nucleic Acids Research (2015) 43(6):3056-3067.
- Silvia Remeseiro and Ana Losada. Cohesin, a chromatin engagement ring. Current Opinion in Cell Biology, (2013) 25(1):63-71.
- Silvia Remeseiro, Cuadrado A, Losada A. Cohesin in development and disease. Development (2013) 140(18):3715-8.
- Silvia Remeseiro, Cuadrado A, Kawauchi S, Calof AL, Lander AD, Losada A. Reduction of Nipbl impairs cohesin loading locally and affects transcription but not cohesion-dependent functions in a mouse model of Cornelia de Lange Syndrome. Biochimica et Biophysica Acta – Molecular Basis of Disease (2013) 3;1832(12):2097-2102.
- Lopez-Contreras AJ, Ruppen I, Nieto-Soler M, Murga M, Rodríguez-Acebes S, Silvia Remeseiro, Rodrigo-Perez S, Rojas AM, Mendez J, Muñoz J, Fernandez-Capetillo O. A proteomic characterization of factors enriched at nascent DNA molecules. Cell Reports (2013) 3(4):1105-16.
- Silvia Remeseiro, Cuadrado A, Carretero M, Martínez P, Drosopoulos WC, Cañamero M, Schildkraut CL, Blasco MA, Losada A. Cohesin-SA1 deficiency drives aneuploidy and tumourigenesis in mice due to impaired replication of telomeres. EMBO Journal (2012) 31:2076-2089.
- Silvia Remeseiro*, Cuadrado A*, Gómez-López G, Pisano DG, Losada A. A unique role of cohesin-SA1 in gene regulation and development. EMBO Journal (2012) 31:2090-2102.
- Cuadrado A, Silvia Remeseiro, Gómez-López G, Pisano DG, Losada A. The specific contributions of cohesin-SA1 to cohesion and gene expression: implications for cancer and development. Cell Cycle (2012) 11(12):2233-8.
- Carretero M, Silvia Remeseiro, Losada A. Cohesin ties up the genome. Current Opinion in Cell Biology (2010) 22(6):781-7.
- Baquero-Montoya C, Gil-Rodríguez M, Teresa-Rodrigo M, Hernández-Marcos M, Bueno-Lozano G, Bueno-Martínez I, Silvia Remeseiro, Fernández-Hernández R, Bassecourt-Serra M, Rodríguez de Alba M, Queralt E, Losada A, Puisac B, Ramos F, Pié J. Could a patient with SMC1A duplication be classified as a human cohesinopathy? Clinical Genetics (2013) May 17. Doi: 10.1111/cge. 12194, 2014:85:446-451.
- Ruiz-Romero C, Carreira V, Rego I, Silvia Remeseiro, López-Armada MJ, Blanco FJ. Proteomic analysis of human osteoarthritic chondrocytes reveals protein changes in stress and glycolysis. Proteomics (2008) 8(3):495-507.
Other publications
- Silvia Remeseiro#. Nya mål för att blockera neuron-till-gliom-synapser i glioblastom. Neurologi i Sverige (2024) nr 1-24.
- Silvia Remeseiro#. Nya mål för att blockera neuron-till-gliom-synapser i glioblastom. Onkologi i Sverige (2024) nr 2-24.

Ten projects at Umeå University receive a total of 30 million SEK from the Swedish Cancer Society.
the 3D organization of DNA can influence the progression of aggressive brain tumour, reveils a new study.
My group is currently searching for new group members, if you would like to know more please do not hesitate to contact me, silvia.remeseiro@umu.se
My group is currently searching for new group members, if you would like to know more please do not hesitate to contact me, silvia.remeseiro@umu.se

Ten projects at Umeå University receive a total of 30 million SEK from the Swedish Cancer Society.
the 3D organization of DNA can influence the progression of aggressive brain tumour, reveils a new study.

25 young researchers are selected to meet to be inspired and inspire others.